E3 Ligase Drug Discovery, Profiling & Screening
Advanced reagents and assays for comprehensive E3 ligase screening and profiling solutions
Need Help?

- Overview
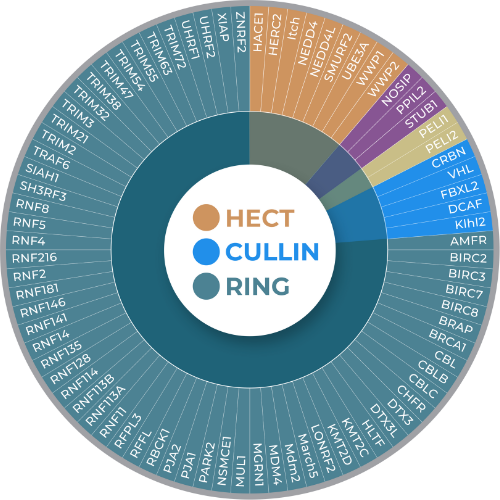
At LifeSensors, we have developed reagents and assays encompassing both traditional and non-traditional approaches for E3 ligase screening and profiling. LifeSensors’ small molecule library contains a collection of ligase-centric compounds, which can be employed as a gold standard for your ligase drug discovery efforts. Further, LifeSensors has the tools to quickly exclude off-target hits through a variety of complementary biochemical and biophysical validation assays. In E3 ligase drug discovery, connecting the dots is particularly important.
Quickly distinguishing true hits from false positives, developing structure-activity relationships, and establishing rank order potency from purified enzymes to cellular models, are key steps to success. LifeSensors brings together the tools and expertise necessary to overcome the many pitfalls in the E3 ligase field.
Understanding E3 Ligase Classifications
Important introduction to E3 Ligases and new classifications beyond the standard familiy format (2025)
E3 Ligase Key Tools for Discovery
HTS Biochemical Assays
LifeSensors offers a range of powerful assays that can be run in-house, from gel and TR-FRET assays, to auto-ubiquitination and substrate ubiquitination assays
Substrate & Ligase Finder and Profiler
Taking advantage of our HTS assays discover how a ligase of interest binds and ubiquitinates a target
Gel-Based Assays
Gel-based ubiquitination assays provide direct, confirmation of ubiquitination and ligase activity, valuable for mechanistic validation in E3 ligase, PROTAC, and molecular glue studies.
Cell-Based Assays
With multiple in-house developed ELISA-based assays, LifeSensors is uniquely suited to evaluate post-translation modifications in intact cells
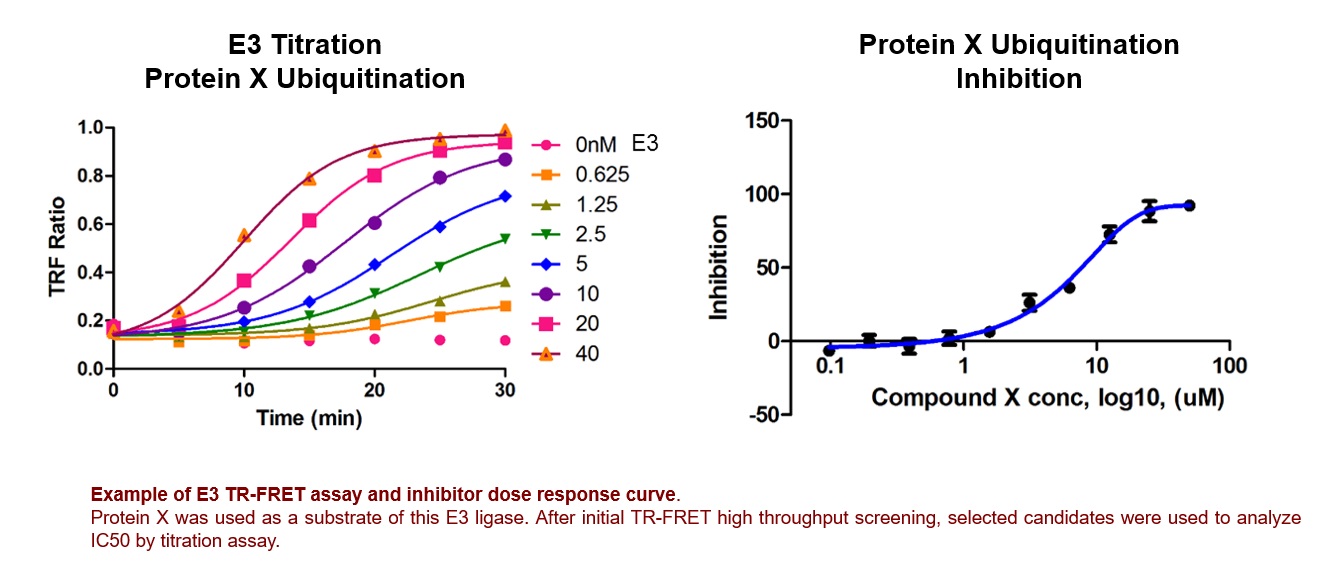
E3 ligases remain among the most difficult enzyme targets to assay and discover potent therapeutic applications. This is possibly due to their dependence on upstream E1 and E2 enzymes for the activation and transfer of ubiquitin. As a result, traditional assays for E3 ligases are complex and prone to off-target false positives. At LifeSensors, we have developed reagents and assays encompassing both traditional and non-traditional approaches for E3 ligase screening and profiling. Quickly distinguishing true hits from false positives, developing structure-activity relationships, and establishing rank order potency from purified enzymes to cellular models, are key steps to success. LifeSensors brings together the tools and expertise necessary to overcome the many pitfalls in the E3 ligase field.
Our E3 TR-FRET detection system utilizes our TUBE technology to monitor E3 ligase activity by fluorescence signal. This assay involves donor-labeled TUBEs that bind to acceptor-labeled polyubiquitin chains synthesized by the target E3 ligase. When donor-labeled TUBEs come in close proximity to polyubiquitin chains containing acceptor-labeled ubiquitin will yield in a FRET signal. This signal can be monitored over time in a homogenous, high-throughput format, making it ideal for small molecule screening to find tool compounds for E3 ligases.
Technology Highlights
- Ubiquitination cascade kits: UC105, UC104
- PROTAC and Molecular Glue Screening kits: PA950, PA770, MG780
- Access to 50+ E3 ligases for performing discovery and selectivity studies in rapid homogenous assay format
- Autoubiquitination and substrate ubiquitination assays: UC109
- Ultrasensitive with robust signal-to-noise powered by TUBE technology
- Extensive experience in UPS related drug discovery – complementary biochemical and biophysical validation studies to exclude off-target hits
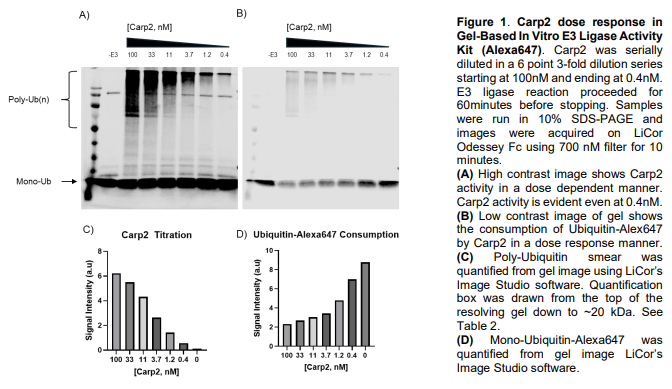
Gel-based assays for evaluating E3 ligases rely on in vitro ubiquitination reactions reconstituted with E1, E2, a chosen E3 ligase, ubiquitin, and ATP, followed by SDS-PAGE and Western blotting to visualize ubiquitination. These assays are commonly used to assess E3 activity, auto-ubiquitination, substrate modification, E2 specificity, and the effects of mutations or inhibitors. Readouts appear as molecular-weight shifts or smears corresponding to mono- or polyubiquitinated species, detected with antibodies against ubiquitin, the E3, or the substrate. While gel-based approaches are low-cost and mechanistically informative, they are semi-quantitative and limited in throughput and chain-type resolution, often serving as a foundation for more advanced biochemical or mass spectrometry-based analyses.
LifeSensors offers a range of technologies and assays for gel-based evaluation, off the shelf, but also provides customer focused development and design in order to assist with project testing.
Technology Highlights
- TUBE-based technology allows for a clear picture of total or lysine specific ubiquitination
- Quantitative measurement utilizing UbiQuant, UbiTest, and other well validated ubiquitin-based assays
- Customizable activity and ubiquitination assays: UC201, UC202
- TUBE-based assays are flexible in gels, western blots, and plate based assays allowing for consistent performance across a variety of platforms
- Autoubiquitination, substrate ubiquitination, and activity assays can deconvolute hits from E3 screens
- Extensive experience in UPS related drug discovery – complementary biochemical and biophysical validation studies to exclude off-target hits from larger screens
In Vitro Ubiquitination
Recreate the ubiquitination cascade to evaluate polyubiquitination with TUBEs. Determine activity and chain length
Substrate Ubiquitination
Take advantage of LIfeSensors protein expression to work with purified substrates and proteins for biologically meaningful data
Custom Assay Design
With many years combined experience working within the Ubiquitin Proteasome System, LifeSensors has the capability to custom design and provide procedural knowledge to take those assays internally
Auto-Ubiquitination
RING and HECT ligases frequently ubiquitinate themselves for a fast readout of E3 catalytic capability. Fast and a great place to start in order to evaluate an E3
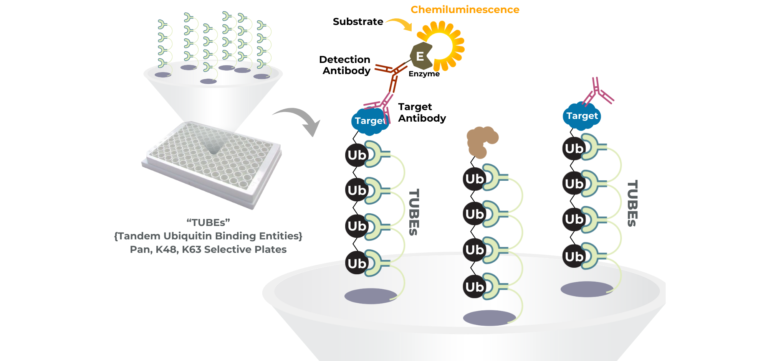
Unlike binding or proximity assays, LifeSensors’ high-throughput screening platforms measure what truly matters for PROTACs and molecular glues: target protein ubiquitination. Powered by proprietary Tandem Ubiquitin Binding Entity (TUBE) technology, these assays capture and quantify polyubiquitinated proteins with exceptional sensitivity and specificity in scalable microplate formats.
TUBE-based HTS assays enable rapid screening and ranking of PROTACs and molecular glues based on their ability to drive ubiquitination through native E3 ligase pathways. Pan- and linkage-selective TUBEs provide additional mechanistic insight by distinguishing degradation-relevant ubiquitin chains (such as K48) from non-degradative signaling events.
By translating ubiquitination biology into a robust, plate-based assay format, LifeSensors’ TUBE-powered HTS platforms help teams move beyond surrogate endpoints and confidently advance the most effective targeted protein degraders.
Technology Highlights
- Well validated assays performed in-house to create a detailed workflow for target exploration
- Off the shelf: PA950, PA770, PA660, PA480, have all been validated as published methods for screening
- Measuring natural ubiquitination via TUBEs avoids artifacts associated with more commonly used tag-based assays leading to better confirmation downstream
- Mechanistic profiling of ubiquitin chain topology
- SAR-driven ranking of PROTAC and molecular glue candidates
- MG780, an in vitro ubiquitination kit, has been developed to establish a high-throughput approach that can accurately predict molecular glue efficiency by monitoring the protein’s intrinsic ability to get ubiquitinated
Chemiluminescent Ubiquitination Assays
Explore the quantitative interaction of a known E3 ligase with unknown substrates, or screen for E3s that target a substrate to determine E3-substrate pairs
TUBE Mass Spec Proteomics
The most detailed evaluation of a target's ubiquitination status coupled with lysine specific TUBEs as a key preparation reagent allowing a high level of detail, important for evaluating lysine specific ubiquitination
Our substrate and E3 finder platform provide a powerful tool for molecular glue discovery by directly measuring E3 ligase activity and substrate ubiquitination in a controlled, quantitative setting. Molecular glues function by stabilizing or inducing interactions between an E3 ligase and a target protein, so detecting enhanced ubiquitination of a candidate substrate is a key readout of glue activity. This platform leverages proprietary polyubiquitin capture reagents to sensitively detect ubiquitination events, allowing researchers to determine whether a small molecule promotes E3-dependent ubiquitin transfer to a target of interest.
The chemiluminescent 96-well plate format enables parallel evaluation of multiple substrates, E3 variants, and compound conditions, making it suitable for screening libraries of candidate molecular glues and quantifying their relative potency. When a high-quality substrate antibody is available, the assay provides a specific, direct readout; when antibodies are lacking, mass spectrometry–based proteomics (via the E3 ID service) can identify novel ubiquitination events induced by candidate compounds. Together, these capabilities allow researchers to identify potential molecular glues, confirm their mechanism of action, and prioritize leads for further biochemical or cellular validation.
Technology Highlights
- A deep exploration of the substrates engaged with a target E3 ligase to identify targets
- Off the shelf: PA950, PA770, PA660, PA480, have all been validated as published methods for identifying E3s
- Assays can be taken in-house to advance the project on additional targets or screen expansion
- Colorimetric readouts improve turnaround time and better quantification relative to western blot measurements
- Substrate ubiquitination, E3 ID, and activity assays, can deconvolute hits from E3 screens
- Flexible investigation based on current research, methodologies, and required level of detail
Chemiluminescent Ubiquitination Assays
Explore the quantitative interaction of a known E3 ligase with unknown substrates, or screen for E3s that target a substrate to determine E3-substrate pairs
TUBE Mass Spec Proteomics
The most detailed evaluation of a target's ubiquitination status coupled with lysine specific TUBEs as a key preparation reagent allowing a high level of detail, important for evaluating lysine specific ubiquitination
LifeSensors offers a multitude of cell-based services in order to convert biochemical findings into a cellular context. TUBEs allow for capture and quantification of ubiquitin in live cells, bacteria, yeast, mammalian, and plant cells, opening the door for drug discovery, target validation, and mechanistic studies without relying entirely on immunoprecipitation and western blots.
These assays are very well validated for over a decade and offer important information regarding ubiquitination, deubiquitinase performance, substrate performance, E3 activity all in response to the introduction of a treatment molecule, PROTAC, molecular glue, or and induced protein stabilizer.
Cellular Assays: an Overview
- PROTAC and molecular glue ubiquitination assays
- Quantitative measurement of ubiquitination utilizing UbiQuant, UbiTest, and other well validated ubiquitin-based assays
- Assays available to be taken in-house to advance the project on additional targets
- Off the shelf: PA950, PA770, PA660, PA480, have all been validated as published methods for screening
- Autoubiquitination, substrate ubiquitination, and activity assays can deconvolute hits from E3 screens
- Extensive experience in UPS related drug discovery – complementary biochemical and biophysical validation studies to exclude off-target hits
Besides aiding drug discovery, E3 ligase screening and profiling services at LifeSensors also offer identification of E2 pairs for E3 ligase of interest, identification of the nature of polyubiquitination on E3 ligase and protein of interest, in order to gain insights into signaling mechanisms using chain selective TUBE technology.
In Vitro Ubiquitination
Recreate the ubiquitination cascade to evaluate polyubiquitination with TUBEs. Determine activity and chain length
Substrate Ubiquitination
Take advantage of LIfeSensors protein expression to work with purified substrates and proteins for biologically meaningful data
Custom Assay Design
With many years combined experience working within the Ubiquitin Proteasome System, LifeSensors has the capability to custom design and provide procedural knowledge to take those assays internally
Auto-Ubiquitination
RING and HECT ligases frequently ubiquitinate themselves for a fast readout of E3 catalytic capability. Fast and a great place to start evaluating E3s
- Resources
Learn More About Our E3 ELISA Assay Here
Slides to learn about our E3 technologies
Learn More About UbiTest Substrate Validation Here
Quantification ubiquitination
HTS Screening of Polyubiquitin Linkage-Specific Molecular Glues
2025 paper describing linkage specific screening for K48 and K63 ubiquitination
Capture and Quantification of Endogenous Polyubiquitinated Proteins From PROTAC treated cells
Download our comprehensive guide
Understanding E3 Ligase Classifications
Important introduction to E3 Ligases and new classifications beyond the standard familiy format (2025)
- Related Products
- E3 Ubiquitin Ligase Assays
Product / Tools Name
SKU
Categories
Add to cart
- Molecular Glue Degraders
