Molecular Glue Degraders
Advancing targeted protein degradation through innovative discovery tools
Need Help?

- Understanding Molecular Glue Degraders
PROTACs have opened new vistas of targeted protein degradation. However, complex medicinal chemistry, warhead optimization, linkerology, and inefficient methods to analyze degradation of the target proteins in cells has hampered its progress. Historical data has shown that optimized molecular glues (MGs) perform the same function as PROTACs.
Molecular glues are a targeted protein degradation modality that operate within the same Ubiquitin Proteasome System as PROTACs but through a distinct, non-bifunctional mechanism. Unlike PROTACs, which are rationally designed heterobifunctional molecules that physically tether a target protein to an E3 ligase, molecular glues are typically monovalent small molecules that induce or stabilize a productive interaction between the two, leading to ubiquitination and proteasomal degradation of the target. These molecules can often be expressed with simple chemistry and LifeSensors provides a plethora of tools to discover and test the performance of molecular glues.
- Key Figures

With deep and diverse expertise in medicinal chemistry, biochemistry, and cell biology specializing in ubiquitin-based drug discovery for the last 15 years, LifeSensors is proud to offer efficient methods to accelerate your discovery of novel molecules facilitating targeted protein degradation.
2
Classes of Glues
E3 ligase binding & target protein binding
- Mechanism of Action
E3 Ligase Binding Class
The first class of glues bind to an E3 ligase to change its conformation so that it binds and ubiquitinates a neo-substrate.
Target Protein Binding Class
The second class of glues bind to a target protein, change its conformation such that the structure is altered, exposing lysines to be ubiquitinated by neighboring ligases.
The net effect is the same – the target protein is ubiquitinated and degraded. Depending upon the type of ligase employed for ubiquitination, if lysine 48 chains are built on the target protein, the proteasome will recognize the target and degrade it. If lysine 63 chains are built on the target protein, the protein may be trafficked to another compartment with a loss of function. However, lysine 63 chains most likely promote autophagy, and the protein will be degraded by lysosomal degradation mechanisms.
LifeSensors has a variety of tools to discover and tease out the mechanism of molecular glue-mediated ubiquitination and degradation.
- Advantages of Choosing LifeSensors For Your Molecular Glue Drug Discovery
- Study molecular glue-mediated ubiquitination and degradation simultaneously
- High-throughput homogenous ubiquitination assays to screen for molecular glues
- Experience with novel E3 ligases ~50 E3 ligases amenable to PROTAC discovery
- Proteomics to characterize protein degradation – establish selectivity
- Our Advanced Methodology
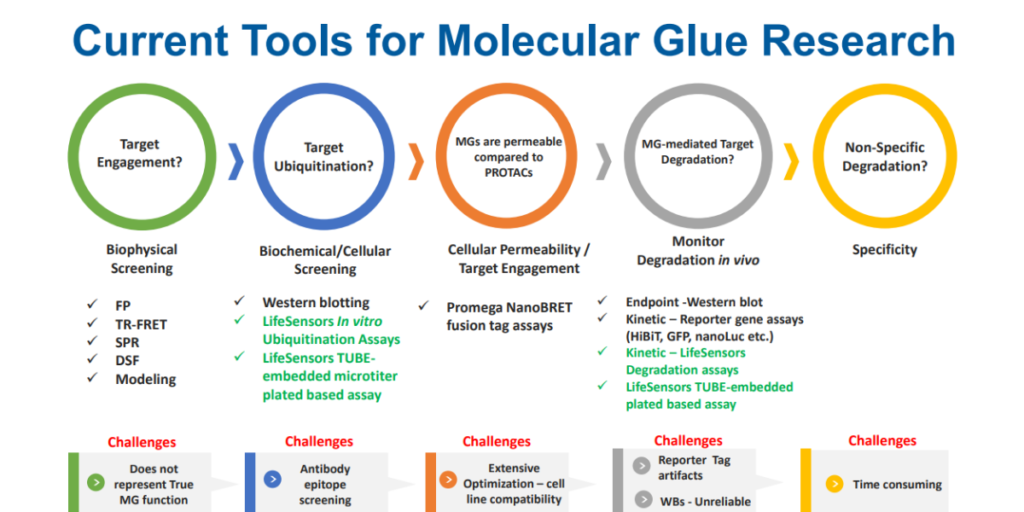
Traditional methods such as western blotting and reporter gene assays have been at the forefront of studying targeted protein degradation. However, these methods are extremely low-throughput, time consuming, and/or prone to artifacts.
LifeSensors’ experience in developing ubiquitination assays for PROTACs allows for that same lysine specific ubiquitination evaluation of molecular glues. Pan-selective Tandem Ubiquitin Binding Entities (TUBEs) still engage with ubiquitin chains regardless of their source, while lysine specific TUBEs can evaluate K48 or K63 ubiquitination evaluate the degradation or trafficking capability of a molecule.
LifeSensors has developed high-throughput methods to directly monitor molecular glue and PROTAC-mediated ubiquitination and degradation of target proteins. Monitoring ligand-mediated ubiquitination of target proteins is the best and the most faithful form of molecular glue or PROTAC function.
- HTS TR-FRET
HTS TR-FRET (High-Throughput Screening Time-Resolved Förster Resonance Energy Transfer) is a biochemical assay format widely used to detect protein–protein interactions, making it particularly well suited for studying molecular glues.
Molecular glues function by stabilizing or inducing an interaction between a target protein and an E3 ligase, often forming a complex that does not exist or is very weak in the absence of the compound. TR-FRET assays can directly measure this induced proximity by labeling the two proteins with a donor and acceptor fluorophore pair. When the proteins come into close proximity, energy transfer occurs, producing a measurable fluorescence signal.
- TR-FRET Key Benefits
Direct readout of protein-protein stabilization
Detection of even weak transient interactions that molecular glues induce without relying on downstream degradation assays
Mix and Read Adapatation
Eliminating steps allows for miniaturization for larger scale screening, and the ability for clients to take this assay type in house for continued work
Time-resolved measurement
Strong donor fluorophores in TR-FRET reduce background autofluorescence to increase assay sensitivity key for detecting subtle effects of molecular glues
LifeSensors TUBE supported HTS TR-FRET allows researchers to identify compounds that stabilize a target E3 ligase complex, rank their relative potencies, and generate SAR (Structure–Activity Relationship) data prior to moving into cell-based ubiquitination or degradation assays.
- Gel-Based Assays
Gel-based assays are a traditional biochemical approach to detect ubiquitination events induced by molecular glues. Typically, the target protein and E3 ligase are incubated with ubiquitin, ATP, and the molecular glue in vitro. Samples are then separated by SDS-PAGE and analyzed by Western blot using antibodies against the target protein or alternatively TUBEs against the ubiquitin chain. Molecular glues that stabilize the target–ligase interaction increase ubiquitination of the target, which appears as a “ubiquitin ladder” or high molecular weight smear on the gel.
LifeSensors’ TUBEs offer further benefit where ubiquitination can be identified as K48, implying likely degradation, or K63, indicating protein trafficking. These assays are low cost and serve as a good initial starting point for evaluating ubiquitination of a target protein in response to the presence of a molecular glue.
While LifeSensors offers a variety of assays off the shelf for measuring ubiquitination, qualitatively as a traditional gel, or quantitatively through chemiluminescent or colorimetric type assays, we are flexible to meet customer design requirements and are interested in assisting companies with their own screening and pipelines.
Service Highlights
- TUBE-based technology allows for a clear picture of total or lysine specific ubiquitination
- Quantitative measurement utilizing UbiQuant, UbiTest and other well validated ubiquitin based assays
- Assays available to be taken in-house to advance the project on additional targets
- Gel-based readouts improve turnaround time relative to western blot measurements
- Autoubiquitination, substrate ubiquitination, activity assays can deconvulate hits from E3 screens
- Extensive experience in UPS related drug discovery – complementary biochemical and biophysical validation studies to exclude off-target hits
- Substrate and E3 Finder
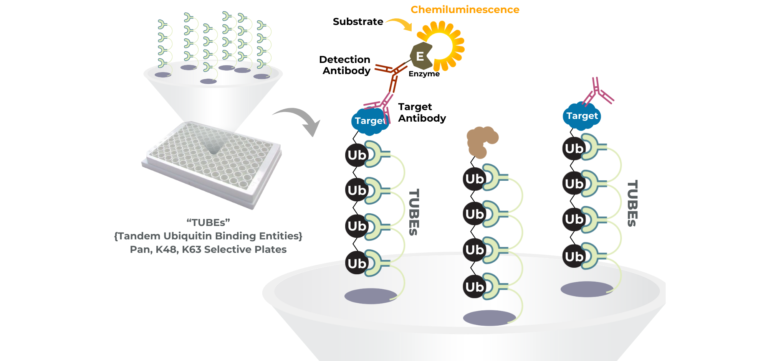
Our substrate and E3 substrate finder platform provide a powerful tool for molecular glue discovery by directly measuring E3 ligase activity and substrate ubiquitination in a controlled, quantitative setting. Molecular glues function by stabilizing or inducing interactions between an E3 ligase and a target protein, so detecting enhanced ubiquitination of a candidate substrate is a key readout of glue activity. This platform leverages proprietary polyubiquitin capture reagents (TUBEs) to sensitively detect ubiquitination events, allowing researchers to determine whether a small molecule promotes E3-dependent ubiquitin transfer to a target of interest.
The chemiluminescent, 96-well plate format enables parallel evaluation of multiple substrates, E3 variants, and compound conditions, making it suitable for screening libraries of candidate molecular glues and quantifying their relative potency. When a high-quality substrate antibody is available, the assay provides a specific, direct readout; when antibodies are lacking, mass spectrometry–based proteomics (via our E3 ID service) can identify novel ubiquitination events induced by candidate compounds. Together, these capabilities allow teams to identify potential molecular glues, confirm their mechanism of action, and prioritize leads for further biochemical or cellular validation. So long as an E3 or substrate is provided, it is possible to find the other.
- Mass Spec Proteomics
Mass Spectrometry (MS) proteomics is a powerful tool for identifying and quantifying ubiquitination events induced by molecular glues. Since molecular glues function by stabilizing or inducing interactions between an E3 ubiquitin ligase and a target protein, they often generate novel ubiquitination patterns on substrates that may not be predicted a priori. MS-based proteomics allows researchers to unbiasedly detect these ubiquitination events across the proteome, capturing both known and previously unrecognized targets of a molecular glue.
In practice, cells or biochemical reactions treated with a candidate molecular glue are lysed, and ubiquitinated proteins are enriched using tools such as TUBEs (Tandem Ubiquitin Binding Entities) or affinity reagents. The enriched proteins are then digested into peptides and analyzed by LC-MS/MS, revealing the specific sites of ubiquitin attachment, the identity of modified proteins, and the relative abundance of ubiquitination. This approach provides mechanistic insight into E3 recruitment and substrate specificity, enabling researchers to validate that a molecular glue induces degradation of the intended target and to identify potential off-target effects.
LifeSensors MS TUBE-based proteomics provides high-resolution, global insight into how small molecules reprogram E3 ligases to selectively ubiquitinate target proteins.
- Ternary Complex Assays
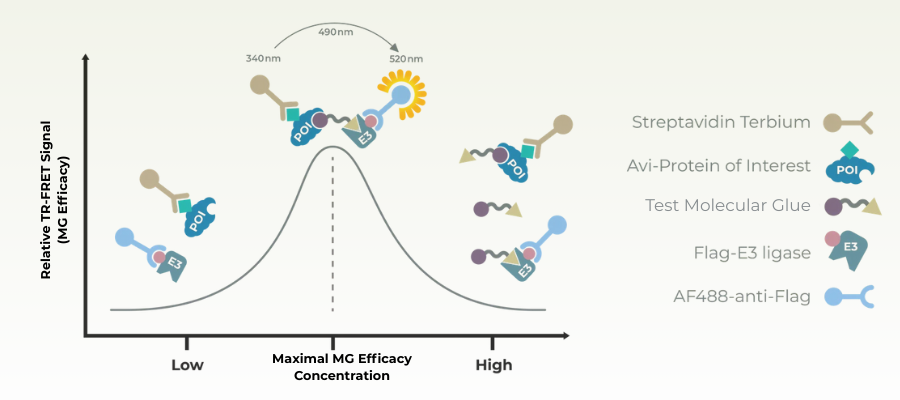
Ternary complex formation assays are a core tool for evaluating molecular glues because they directly measure the induced interaction between a target protein and an E3 ligase, which is the defining mechanism of glue activity. In the absence of a molecular glue, these protein–protein interactions are often weak or nonexistent; a productive glue stabilizes the ternary complex long enough to enable ubiquitination. Ternary complex assays therefore provide an early, mechanism-focused readout that sits upstream of ubiquitination and degradation.
Formation of the ternary complex in the presence of a molecular glue brings the labels into close proximity, generating a measurable signal that correlates with complex stability and compound potency. These assays are well suited for high-throughput screening and SAR development, allowing rapid ranking of compounds based on their ability to promote E3 substrate engagement.
For molecular glue discovery, ternary complex assays are especially valuable because they can distinguish true glue activity from nonspecific effects on ubiquitination or protein abundance. When combined with downstream ubiquitination assays, TUBEs, or proteomics, they help establish a clear mechanistic chain from compound to degradation, reducing false positives and accelerating confident lead selection.
Key Insight:
Studies in recent years have shown that protein degradability is strongly influenced by intrinsic characteristics of the protein, especially the protein’s endogenous “Ubiquitination Potential”. Hence, correlating ternary complex formation with ubiquitination potential is crucial in accurately characterizing molecular glue efficiency and helping select the best candidates as potential drugs.
- Related Products
- Molecular Glue Degraders
