Tandem Ubiquitin Binding Entities (TUBEs) are powerful reagents for enrichment of polyubiquitylated proteins. At LifeSensors, we have used this remarkable technology of TUBEs with our mass spectrometry expertise to provide the customer with a quick and easy way to perform both qualitative and quantitative proteomics. Upon completion of the analysis, you will receive a list of peptides and corresponding proteins that have been identified. We will categorize the ubiquitylated proteins and display the results on several plots to make your analysis easier. This information will help you publish your results or plan your next assay.
We are here to help you from the beginning, and are happy to sit down and help you design the best experiment to address your specific questions. We have competitively priced our analysis and offer custom services with the level of analysis that matches your needs and budget. We strive to deliver you accurate and professional reports quickly. From submission of samples reports are typically delivered in 2-4 weeks depending on the complexity of project. Please contact us for a more accurate timeline for your specific experiment.
TUBE-based enrichment of polyubiquitylated proteins has proven key for the progress of ubiquitin proteomics. LifeSensors has developed the proprietary technology to identify cellular proteins that are ubiquitylated using TUBE-based proteomics. A brief overview of this technology is depicted in the following figure:
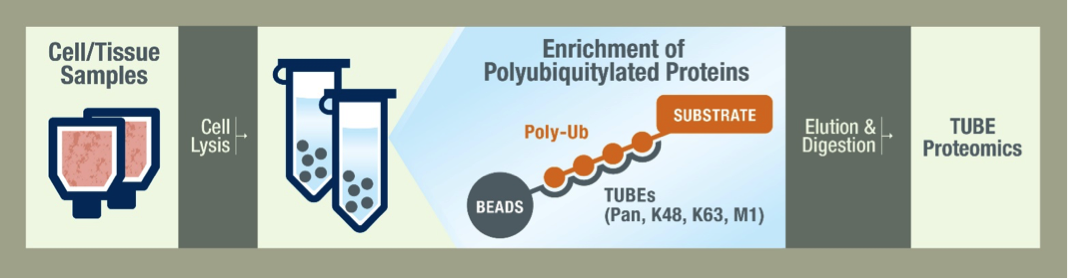
Workflow schematic of TUBE proteomics. Treated cells are enriched using TUBEs (TUBE1 and K48). The enriched polyubiquitylated proteins are pixelated by SDS-PAGE and digested with trypsin. Individual ubiquitylated proteins are identified by the presence of at least 2 peptides in at least 2 of 3 replicates. Ub linkage types are identified by K-ε-GG peptides corresponding to covalent Ub-Ub modification at a particular lysine.
LifeSensors’ pan (TUBE1 and TUBE2) as well as linkage-selective (M1-, K48-, and K63- specific) TUBEs provide isolation of respective polyubiquitylated proteins. The TUBE enrichment kit contains solutions to provide simple procedure to lyse cells, wash the non-specific binding proteins, and elute proteins in solution compatible for further protease digestion. TUBE proteomics services include following four steps:
Chain Selectivity-
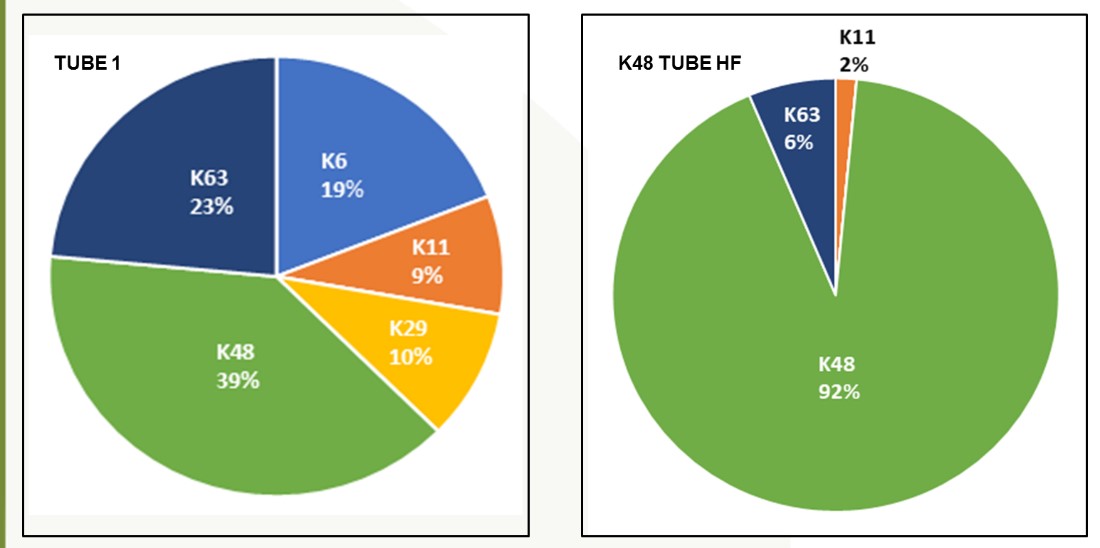
EXPERIMENTAL DATA
In-house data
Below are examples of data generated using TUBE- based mass spec technology with pan- selective TUBEs. Here the ubiquitinated proteins have been differentiated from whole cell lysate. This methodology is sensitive enough to detect the number of ubiquitin sites on a given protein. The number of ubiquitinated proteins detected was assigned to the number of ubiquitin sites that they carry. In this case over half of the ubiquitnated proteins detected were monoubiquitinated. When studying the effects of a drug or genetic knockout global shifts in the ubiquitome is a good place to start your analysis.
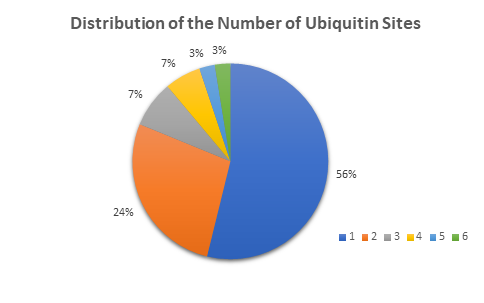
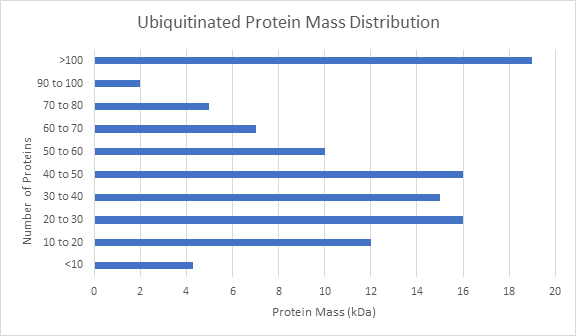
Other global analysis provided by LifeSensors includes protein mass distribution of all proteins as well as ubiquitinated proteins. The number of ubiquitinated proteins detected was assigned to different intervals of protein mass. Here there is an interesting cluster of proteins around 30 kDa, and over 100 kDa. The ubiquitin pathway is clearly playing a role in clearing and otherwise influencing larger proteins in this experiment.
PUBLISHED DATA
Silva et al.1 have used linkage-specific TUBE-based Mass Spectrometry to identify >100 novel K63 polyubiquitylated targets that were significantly enriched in ribosomal proteins. Using this and other data, they show that oxidative stress response is modulated by K63-linked polyubiquitylation.
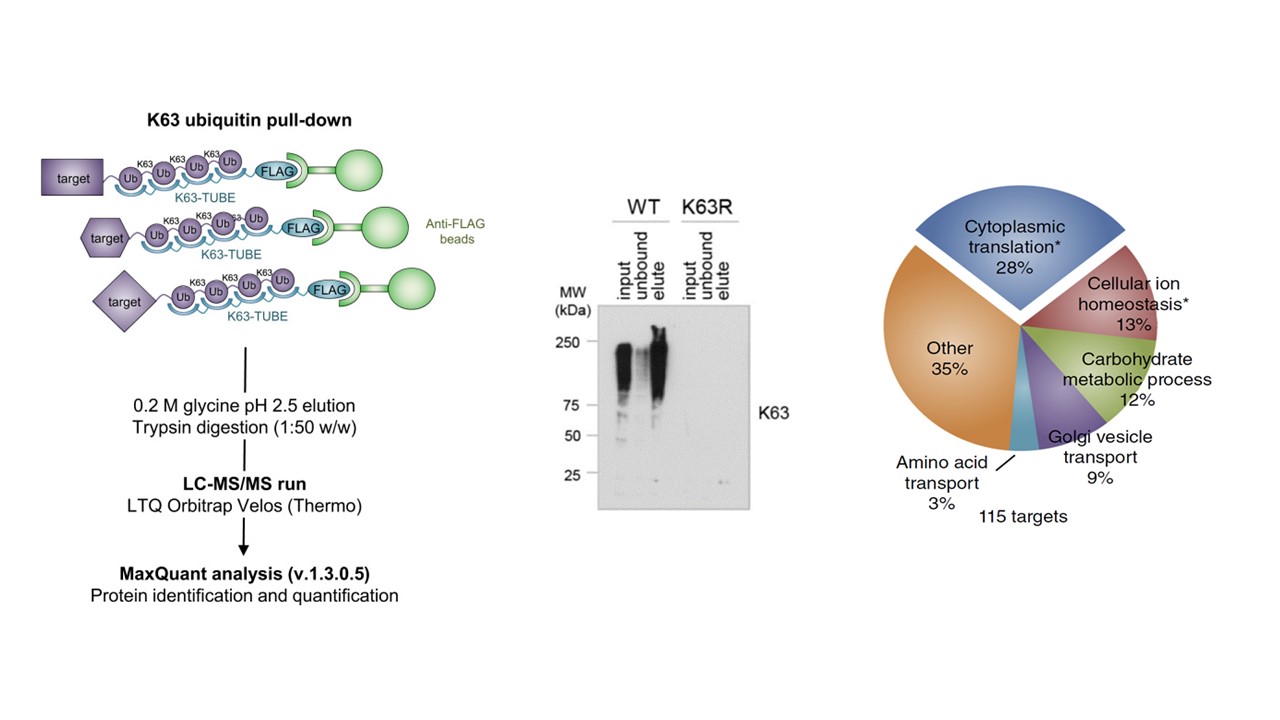
In 2016, Mata-Cantero et al.2 used TUBE-based mass-spectrometry to identify major components of the ubiquitin proteome of both Plasmodium falciparum and its host during different life stages.
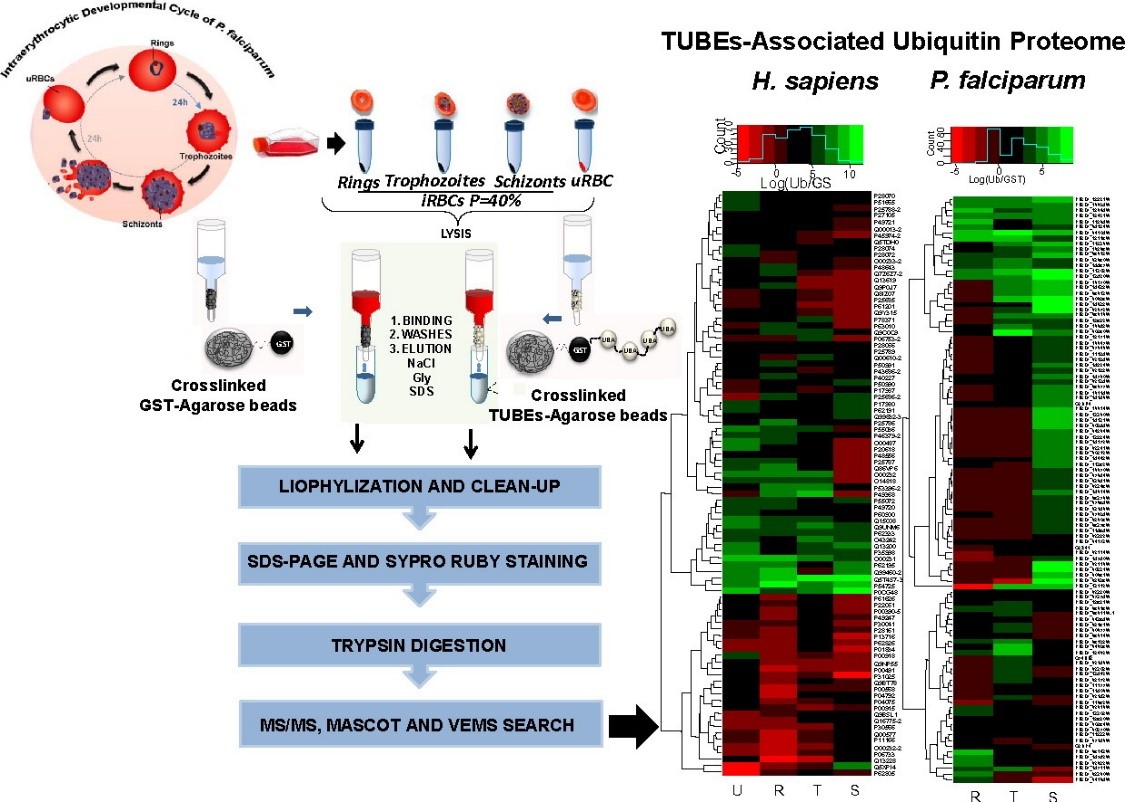
Identification of ubiquitylated proteins using TUBEs-LC-MS/MS method. Intraerythrocytic Developmental Cycle of P. falciparum is shown. Synchronized P. falciparum iRBC at 40% parasitaemia from rings, trophozoites and schizonts stages were collected and frozen. TUBE enriched proteins from iRBC at different stages and uRBC were captured using TUBEs or GST (control) previously crosslinked with DMP to agarose beads. After exhaustive washes, proteins captured were eluted, cleaned by precipitation and resolved by electrophoresis (PAGE). Bands with proteins were analyzed by LC-MS/MS.
References
- Silvaet al. K63 polyubiquitination is a new modulator of the oxidative stress response Nature Structural Molecular Biology 22 (2015) 116–123
- L. Mata-Canteroet al., New insights into host-parasite ubiquitin proteome dynamics in P. falciparum infected red blood cells using a TUBEs-MS approach, Journal of Proteomics, 139 (2016) 45–59
