The Ubiquitin Proteasome System (UPS) controls a majority of the principal functions of almost all the cellular proteins of human cells. Failures in this system often contribute directly or indirectly to the pathogenesis of many diseases, including cancer, inflammation, and neurodegeneration. There are ~700 E3 ubiquitin ligases encoded in human genome and the functions of most of these E3s remain unknown. Recent development of PROTAC drugs to recruit E3 ligases and degrade therapeutic proteins highlight the value of studying E3 ligases. Substrate identification in individual E3 ligases is an essential step to unraveling cellular functions of E3s and identifying novel therapeutic targets. Lifesensors can help you identify the substrates of your preferred E3s.
Service Highlights
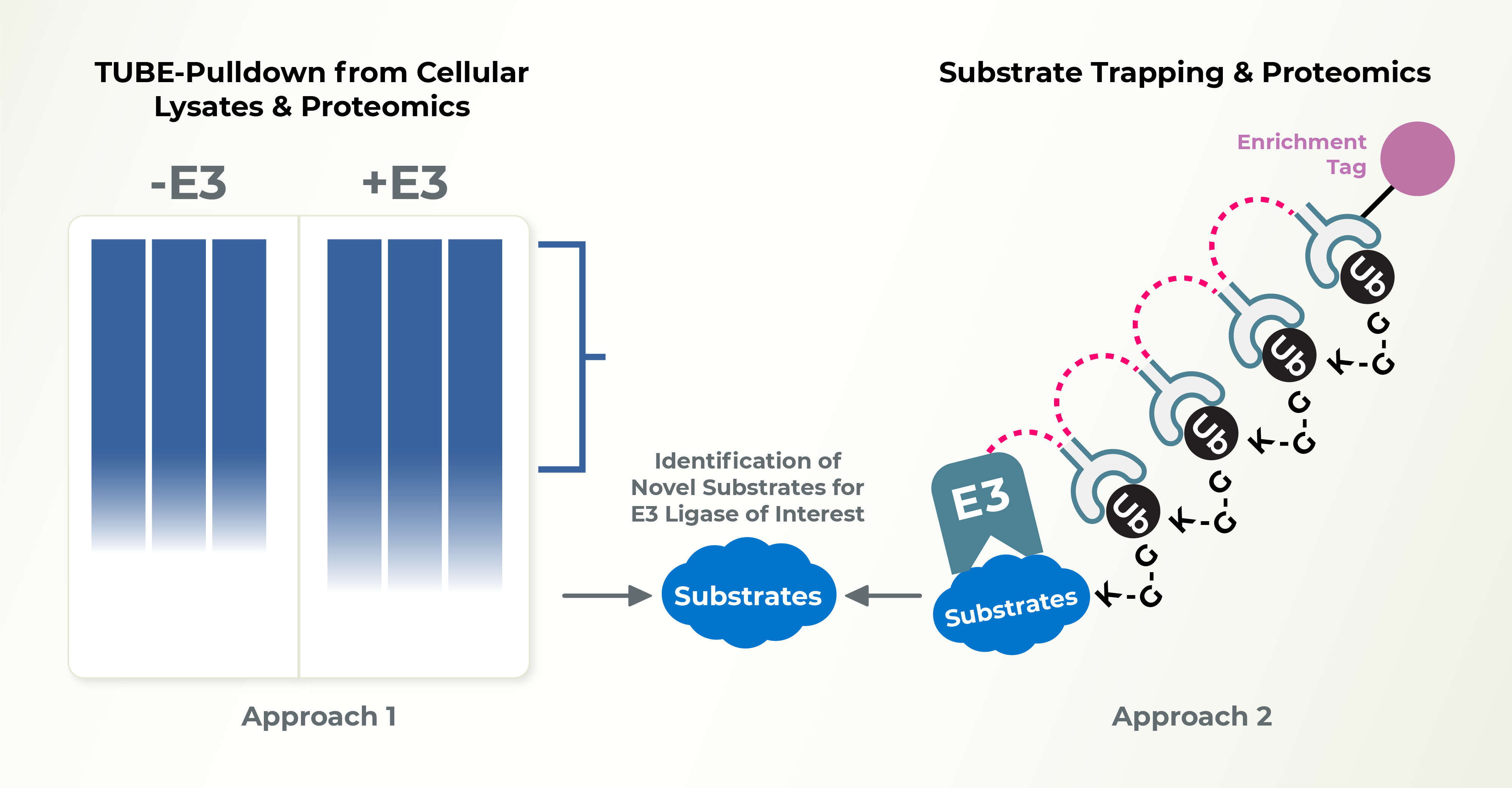
- Cell-based assays to identify endogenous E3 ligase substrates
- With over expression of E3s and/or Knockout/Knockdown of E3s
- Substrate Trapping and Proteomic analysis to identify endogenous substrates
- Validation of substrates in orthogonal assays
Related Products
TUBE1
Coated polymeric high-capacity magnetic beads to allow superior enrichment of polyubiquitinated proteins along with minimizing non-specific binding to proteins in tissue and cellular lysates.
Agarose-TUBE1
TUBEs display up to a 1000-fold increase in affinity for polyubiquitin moieties over the single ubiquitin binding associated domain (UBA). In addition, TUBEs protect polyubiquitinated proteins from deubiquitination and proteasomal degradation, allowing for detection at relatively low abundance.
VU1
Monoclonal antibodies that recognize polyubiquitylated proteins and free ubiquitin. VU-1 has been shown to recognize all ubiquitin linkages (mono, K6, K11, K27, K29, K33, K48, K63, and linear) by Western blotting. VU-1 is an excellent antibody for immunostaining applications generating robust ubiquitin detection and low background with an ability to detect subcellular ubiquitin localization.