LifeSensors recently presented the work of Rajesh Singh, Cari Sagum, Jianji Chen, Apurva Chaturvedi, Peter Foote, and Mark Bedford at Cold Spring Harbor Laboratories. A PDF of this poster can be downloaded Here.
ABSTRACT
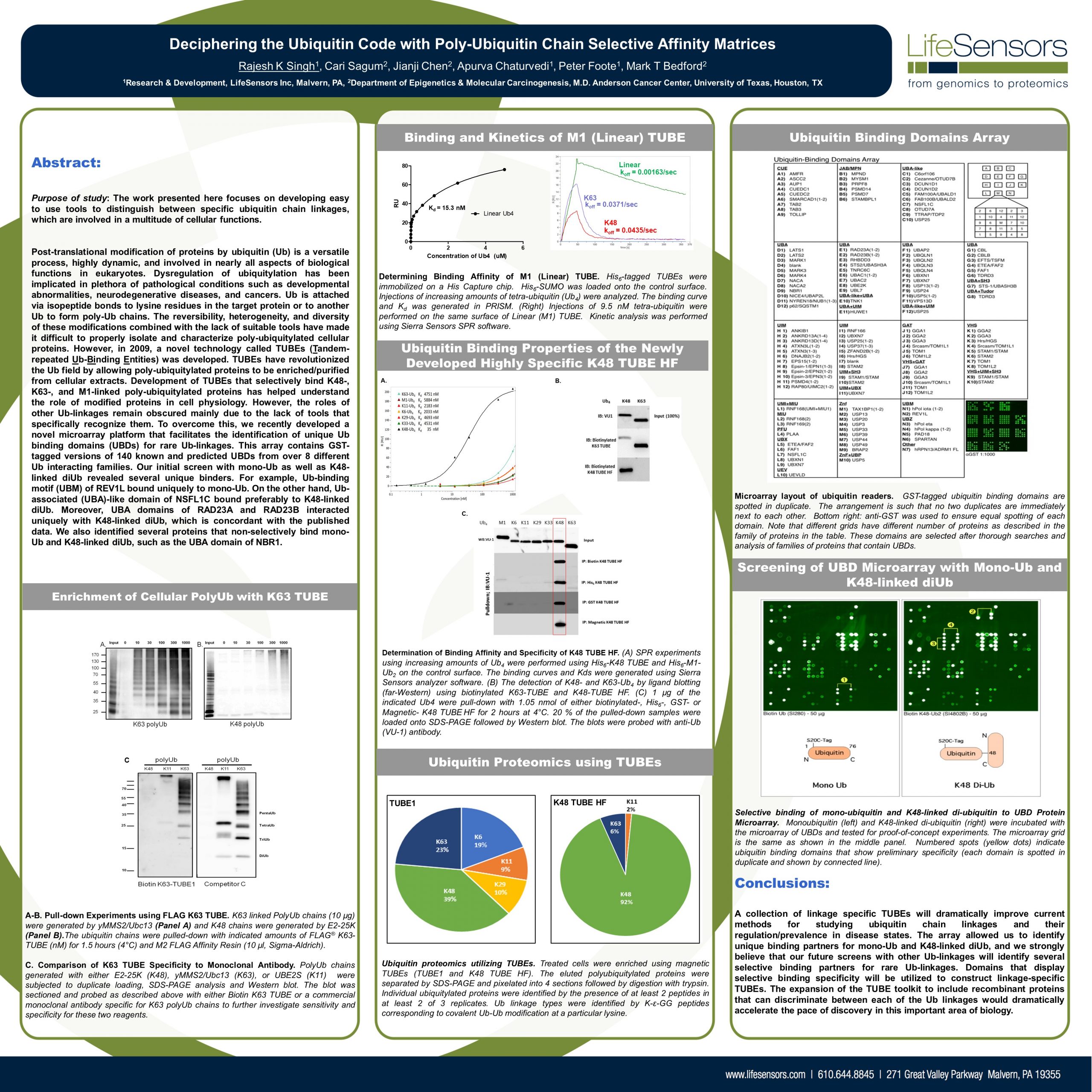
Post-translational modification of proteins by ubiquitin (Ub) is a versatile process, highly dynamic, and involved in nearly all aspects of biological functions in eukaryotes. Dysregulation of ubiquitylation has been implicated in plethora of pathological conditions such as developmental abnormalities, neurodegenerative diseases, and cancers. Ub is attached via isopeptide bonds to lysine residues in the target protein or to another Ub to form poly-Ub chains. The reversibility, heterogeneity, and diversity of these modifications combined with the lack of suitable tools have made it difficult to properly isolate and characterize poly-ubiquitylated cellular proteins. However, in 2009, a novel technology called TUBEs (Tandemrepeated Ub-Binding Entities) was developed. TUBEs have revolutionized the Ub field by allowing poly-ubiquitylated proteins to be enriched/purified from cellular extracts. Development of TUBEs that selectively bind K48-, K63-, and M1-linked poly-ubiquitylated proteins has helped understand the role of modified proteins in cell physiology. However, the roles of other Ub-linkages remain obscured mainly due to the lack of tools that specifically recognize them. To overcome this, we recently developed a novel microarray platform that facilitates the identification of unique Ub binding domains (UBDs) for rare Ub-linkages. This array contains GSTtagged versions of 140 known and predicted UBDs from over 8 different Ub interacting families. Our initial screen with mono-Ub as well as K48-linked diUb revealed several unique binders. For example, Ub-binding motif (UBM) of REV1L bound uniquely to mono-Ub. On the other hand, Ubassociated (UBA)-like domain of NSFL1C bound preferably to K48-linked diUb. Moreover, UBA domains of RAD23A and RAD23B interacted uniquely with K48-linked diUb, which is concordant with the published data. We also identified several proteins that non-selectively.
