LifeSensors’ Scientists Apurva Chaturvedi, Peter Foote, Dominic Vasturia, and Rajesh Singh recently presented their research at AACR2018. A pdf of this poster can be downloaded here.
ABSTRACT
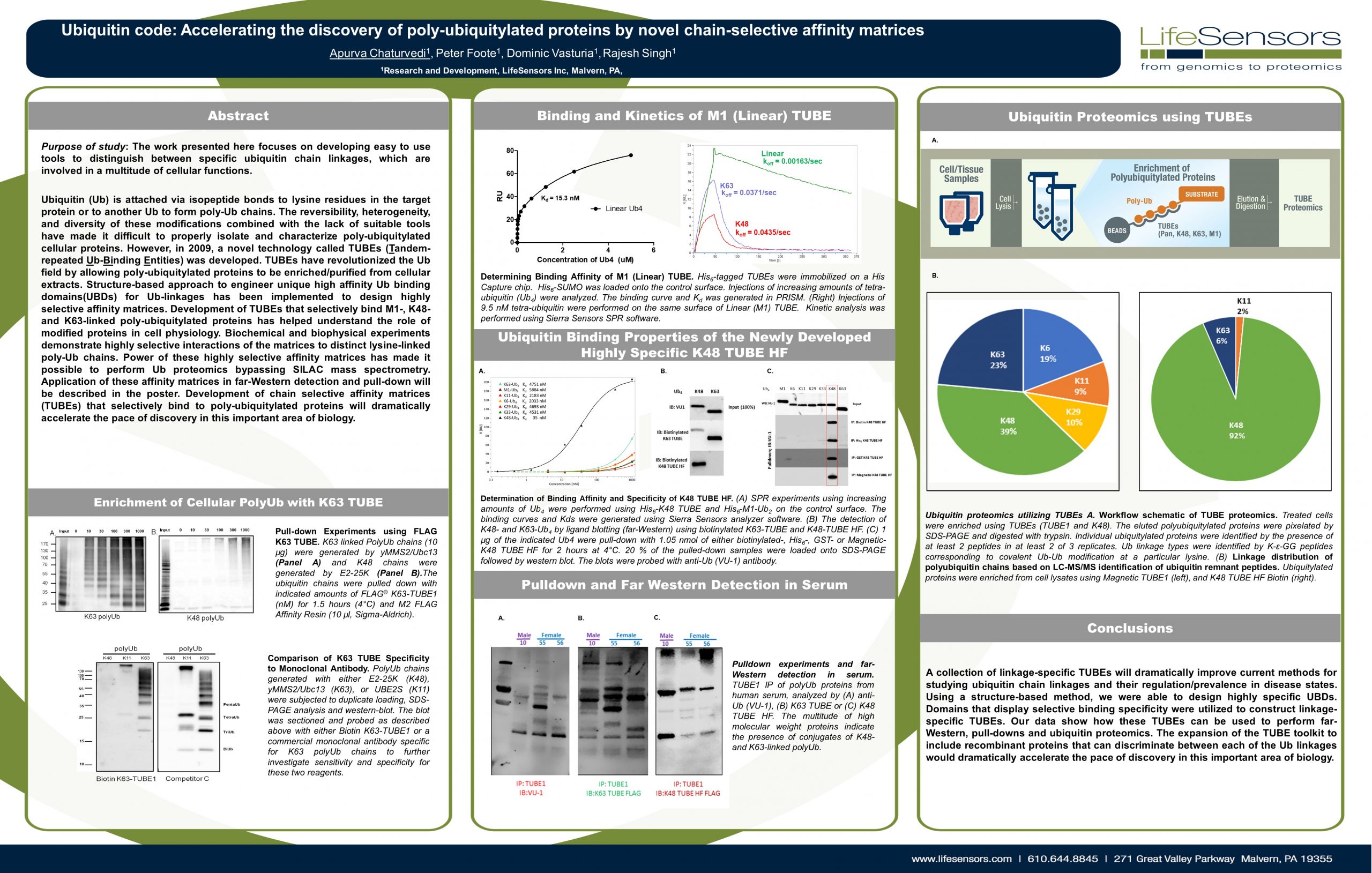
Purpose of study: The work presented here focuses on developing easy to use tools to distinguish between specific ubiquitin chain linkages, which are involved in a multitude of cellular functions.
Ubiquitin (Ub) is attached via isopeptide bonds to lysine residues in the target protein or to another Ub to form poly-Ub chains. The reversibility, heterogeneity, and diversity of these modifications combined with the lack of suitable tools have made it difficult to properly isolate and characterize poly-ubiquitylated cellular proteins. However, in 2009, a novel technology called TUBEs (Tandem-repeated Ub-Binding Entities) was developed. TUBEs have revolutionized the Ub field by allowing poly-ubiquitylated proteins to be enriched/purified from cellular extracts. Structure-based approach to engineer unique high affinity Ub binding domains(UBDs) for Ub-linkages has been implemented to design highly selective affinity matrices. Development of TUBEs that selectively bind M1-, K48-and K63-linked poly-ubiquitylated proteins has helped understand the role of modified proteins in cell physiology. Biochemical and biophysical experiments demonstrate highly selective interactions of the matrices to distinct lysine-linked poly-Ub chains. Power of these highly selective affinity matrices has made it possible to perform Ub proteomics bypassing SILAC mass spectrometry. Application of these affinity matrices in far-Western detection and pull-down will be described in the poster. Development of chain selective affinity matrices (TUBEs) that selectively bind to poly-ubiquitylated proteins will dramatically accelerate the pace of discovery in this important area of biology.
Learn more about TUBEs
