Lifesensors recently presented the work of Peter Foote, Xialong Lu, Apurva Chaturvedi, and Rajesh Singh at AACR2018. A PDF of this poster can be downloaded here.
ABSTRACT
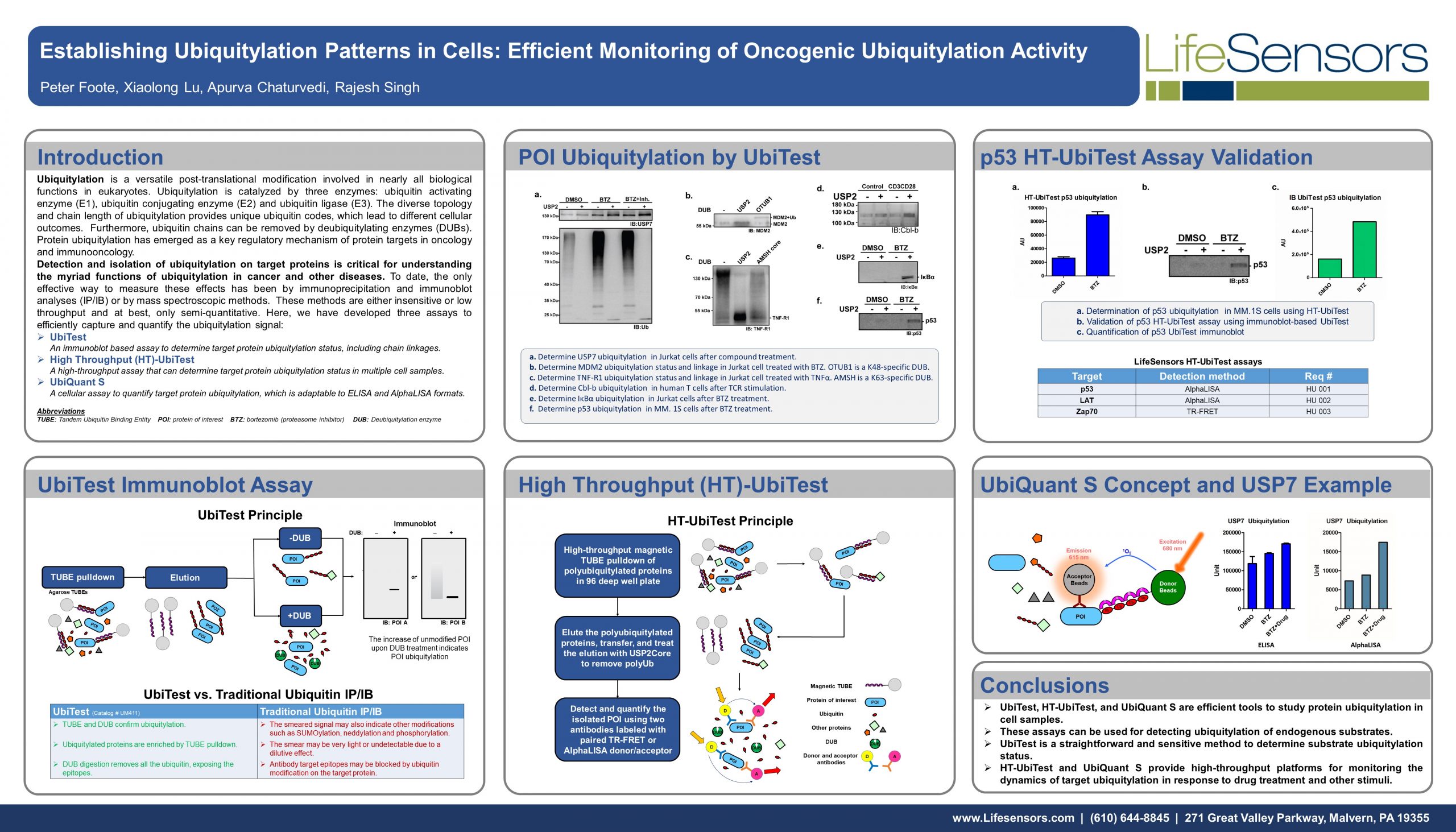
Ubiquitylation is a versatile post-translational modification involved in nearly all biological functions in eukaryotes. Ubiquitylation is catalyzed by three enzymes: ubiquitin activating enzyme (E1), ubiquitin conjugating enzyme (E2) and ubiquitin ligase (E3). The diverse topology and chain length of ubiquitylation provides unique ubiquitin codes, which lead to different cellular outcomes. Furthermore, ubiquitin chains can be removed by deubiquitylating enzymes (DUBs). Protein ubiquitylation has emerged as a key regulatory mechanism of protein targets in oncology and immunooncology.
Detection and isolation of ubiquitylation on target proteins is critical for understanding the myriad functions of ubiquitylation in cancer and other diseases. To date, the only effective way to measure these effects has been by immunoprecipitation and immunoblot analyses (IP/IB) or by mass spectroscopic methods. These methods are either insensitive or low throughput and at best, only semi-quantitative. Here, we have developed three assays to efficiently capture and quantify the ubiquitylation signal:
UbiTest
An immunoblot based assay to determine target protein ubiquitylation status, including chain linkages.
High Throughput (HT)-UbiTest
A high-throughput assay that can determine target protein ubiquitylation status in multiple cell samples.
UbiQuant S
A cellular assay to quantify target protein ubiquitylation, which is adaptable to ELISA and AlphaLISA formats.
